Why Public Health Depends Upon Antimicrobial Stewardship
Antimicrobial resistance (AMR) is an urgent global health threat. A 2022 report published in The Lancet, estimates that 4.95 million deaths were associated with bacterial AMR globally in 2019. In the U.S., more than 2.8 million antimicrobial-resistant infections occur each year. More than 35,000 people die as a result, according to CDC’s 2019 Antibiotic Resistance (AR) Threats Report. According to a CDC study, the estimated national cost to treat infections caused by six multidrug-resistant germs frequently found in health care can be substantial—more than $4.6 billion annually. As hundreds of thousands of patients come face-to-face with health issues related to antibiotic-resistant infections, the scientific community must address the significance of antimicrobial stewardship and its core principles.
What is antimicrobial stewardship?
According to the Centers for Disease Control and Prevention (CDC), antimicrobial stewardship is the endeavor to measure and improve how antibiotics are prescribed and used in a healthcare setting.1 This type of conservancy creates an innovative approach to identifying the critical and complex need for antibiotic therapies, while measuring the outcome to prevent misuse or overuse of antibiotics.
While antibiotics are prescribed to fight bacterial infections, the CDC reports that approximately 30% of all antibiotics prescribed to acute care patients in the United States are not necessary in the progression of treatment.2 Overuse of antibiotics has led to an increase in antimicrobial resistance, which has been identified as a serious ongoing threat to public health.
Who was Sir Alexander Fleming, and how did he contribute to antimicrobial stewardship?
In 1928, Scottish scientist Sir Alexander Fleming made his monumental discovery of penicillin while experimenting with staphylococcal bacteria.3 Quite by accident, he found one of his uncovered agar plates had become contaminated with mold spores, and bacteria surrounding the mold growth were inhibited. He later identified the mold as a member of the Penicillium genus and began to perform further experiments to determine the inhibitory agent. The agent, now termed penicillin, was later found to fight against bacteria that cause diseases such as pneumonia, meningitis and scarlet fever.3
Fleming’s findings did not catch on quickly, as it took more than a decade from the time of his publications for the scientific community to embrace penicillin. It was scientists Howard Florey and Ernst Chain who initiated the use penicillin as a treatment for soldiers during World War II.3 After mass production and successful use of penicillin throughout the war, Fleming was ultimately awarded the Nobel Prize for Physiology/Medicine in 1945.3
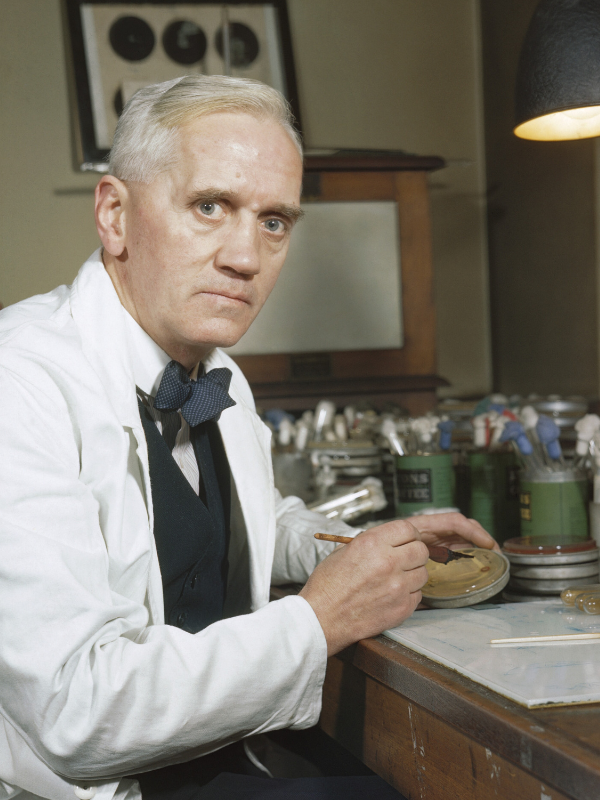

What we now refer to as antimicrobial resistance was first addressed by Fleming during a 1945 interview with the New York Times. While noting the importance of an antibiotic, such as penicillin, he also warned about the abuse of such a critical discovery, “In such cases, the thoughtless person playing with penicillin is morally responsible for the death of the man who finally succumbs to infection with the penicillin-resistant organism. I hope this evil can be averted.”4 As if on cue from Fleming’s ominous prediction, just ten years into its widespread use, penicillin resistance began to emerge.
Rising resistance
During the past few decades, many strains of bacteria have evolved resistance to antibiotics. Infectious bacteria are much more difficult to control than their predecessors were ten or twenty years ago.
The uptick in multi-drug resistant organisms (or MDROs) has resulted in the magnification of hospital-acquired infections (HAI) and a decrease in antibiotic efficacy against certain microorganisms. These consequences bring complexities to medical decisions regarding patient treatment and patient management. MDRO diagnosis in a hospital or long-term care setting requires additional controls, patient isolation, often-prolonged patient hospitalization, and the use of last-line-of-defense treatments, which bring even further risk.
While the pace of antibiotic discovery was stunted in the early 2000s, there has been an uptick of new broad-spectrum antibiotics approved by the FDA within the last ten years geared toward MDROs such as Ceftolozane/Tazobactam, Ceftazidime/Avibactam, Meropenem/Vaborbactam, Delafloxacin, Plazomycin, Omadacycline, Eravacycline, Lefamulin, Imipenem/Relebactam, and Cefiderocol. In order to confront the matter of the escalation of antimicrobial resistance, hospitals and medical staff must establish procedures in correlation to an antimicrobial stewardship mindset.
How hospitals and clinicians can help
Saving patients from contracting resistant organisms must start with initiating the development of an antimicrobial stewardship group within the facility. There are several elements needed for an effective infection prevention team: leadership, accountability, drug expertise, action, tracking, reporting and education.7 These key points encompass the mission of antimicrobial stewardship.

Leadership comes first in the medical institution by simply incorporating an antimicrobial stewardship standard of duty in professional job descriptions, exhaustive stewardship training, and contractual obligations required by those positions. With that principle at the foundation, accountability follows by forming policies to support antibiotic use, requiring interventions with certain prescriptions, and reassessment by a subject matter expert (i.e. pharmacists, clinicians, etc.)
Second, the use of broad-spectrum antibiotics should be discouraged in favor of more narrow spectrum antibiotics that can be directed only to the pathogenic organism.
Third, the use of empirical therapy, which guesses at the appropriate antimicrobial drug, should be discouraged in favor of performing Antimicrobial Susceptibility Tests (AST) in order to select the most effective drug for better patient outcomes.
All of these steps ultimately help achieve a successful antimicrobial stewardship program, which is the core consensus of the process.
Hardy Diagnostics' role in the fight against antimicrobial resistance
Patient health is our priority. We know antimicrobial resistance is on the rise. We also know that our company can and has made a difference by focusing on detection, identification and diagnosis to assist laboratorians and physicians in keeping patients safe and healthy. Antimicrobial stewardship and reducing spread of resistant genes to pathogenic bacteria by detection of these resistant organisms is key to this fight.
Hardy offers an array of products focused on promoting antimicrobial stewardship, from environmental monitoring to patient management, including culture media, antimicrobial diffusion disks, and rapid identification tests targeted at rapid detection of multi-drug resistant organisms.
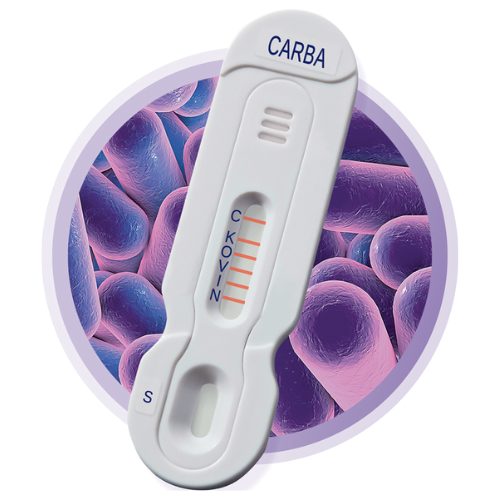
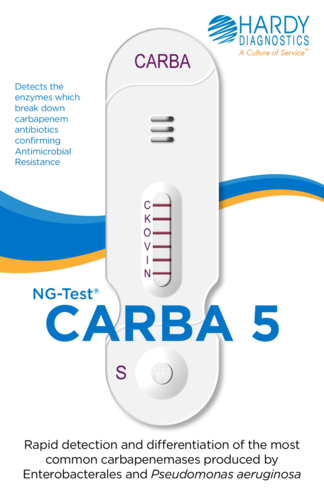
NG-Test® CARBA 5
Carbapenem antibiotics are one of the few remaining antibiotics effective in treating ESBL infections or MDRO.9 However, carbapenemases, or enzymes produced by bacteria that cause resistance to this class of antibiotics, are also on the rise. NG-Test® CARBA 5 is the only FDA cleared, in vitro, rapid, multiplex, phenotypic lateral flow confirmatory test for the qualitative detection of five common carbapenemases produced by Enterobacterales and Pseudomonas aeruginosa: KPC, OXA-48-like, VIM, IMP, and NDM. This lateral flow test can be used with colonies that screen positive via HardyCHROM™ CRE or colonies from blood or MacConkey agar. CARBA 5 is a game-changer in effective patient management, allowing clinicians a clear and rapid understanding of which drugs will benefit patient treatment from those that will not. This allows for better outcomes and more cost-effective care, better control over potential HAI, and aids in the furtherance of antimicrobial stewardship.
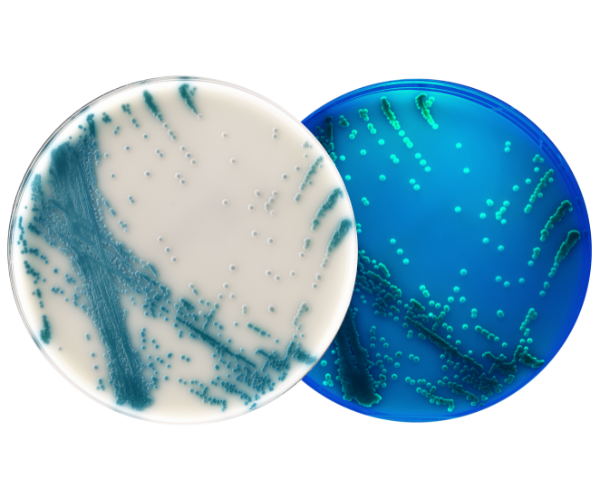
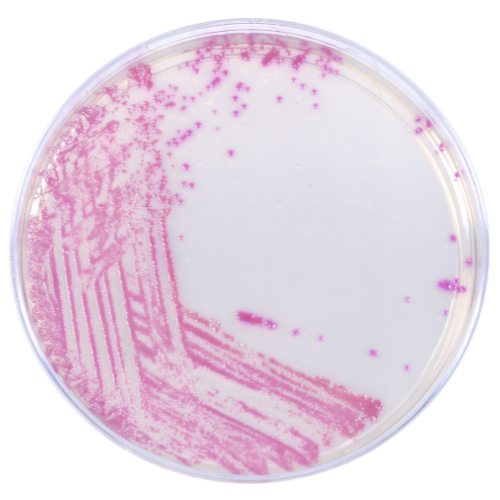
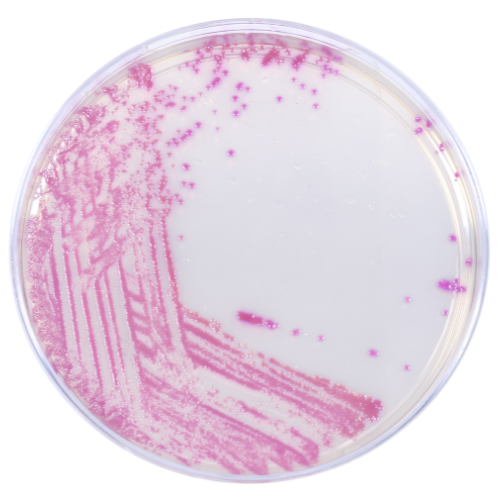
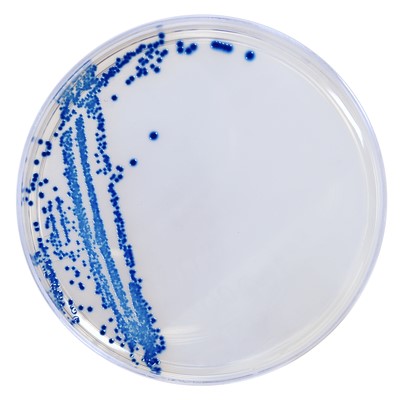
HardyCHROM™ CRE
HardyCHROM™ CRE (Carbapenem Resistant Enterobacterales) is a selective and differential chromogenic agar medium intended for use with stool specimens for the qualitative and presumptive detection of Escherichia coli that are non-susceptible to carbapenems as pink colonies and KES (Klebsiella aerogenes, Klebsiella oxytoca, Klebsiella pneumoniae, Enterobacter cloacae complex, and Serratia marcescens) that are non-susceptible to carbapenems as blue colonies.
HardyCHROM™ Candida + auris
HardyCHROM™ Candida + auris is recommended for the selective isolation and differential indentification of Candida species. The medium allows for the differentiation of Candida albicans, Candida tropicalis, Candida krusei, and Candida auris based on colony morphology, color, and positive UV fluorescence.
Candida auris produces white colonies with teal to teal-green “bullseye” centers that are positive for UV fluorescence for easy differentiation from other Candida species.
HardyCHROM™ MRSA
Studies show 33% of the population carry S. aureus as part of their microbiome. Furthermore, the CDC estimates two out of every 100 people routinely carry MRSA, most commonly in their anterior nares.8 Though most do not develop a serious infection, the potential to transmit MRSA to other patients in a healthcare setting is significant. Over the past five decades, the CDC has engaged in surveillance of HAI using a combination of voluntary reporting and a national system of sentinel hospitals. HardyCHROM™ MRSA can be used in that capacity, and is recommended for the qualitative detection of nasal colonization by MRSA to aid in the prevention and control of this miorganism.
HardyCHROM™ ESBL
Regarding the Gram negative organisms, ESBLs are enzymes produced by certain members of Enterobacterales that can break down and destroy commonly used antibiotics, such as broad spectrum penicillins or cephalosporins, making these drugs ineffective in treating infections. These mechanisms of resistance have existed for some time, first emerging in the medical literature as a β-lactamase (narrow spectrum) identified in Escherichia coli even prior to the medical use of penicillin.10 However, the number of known β-lactamases has quickly grown and, being plasmid and transposon mediated, they’ve spread rapidly world-wide. These mechanisms of resistance have broadened to new enzymes capable of hydrolyzing newer β-lactam antibiotics, leaving very few remaining effective antibiotic treatment options available for patient care. HardyCHROM™ ESBL can aid in the detection of these microorganisms. This medium is intended for the qualitative and presumptive detection of Enterobacterales potentially non-susceptible to broad spectrum cephalosporins, and ESBL-producing E. coli, Klebsiella pneumonia, and K. oxytoca from stool.
Available support for clinical laboratories

Here at Hardy Diagnostics, we take our mission statement to heart: “We are committed to producing and distributing the finest products for the detection of microorganisms, as we partner with our laboratory customers to diagnose and prevent disease.” Hardy Diagnostics pledges to continue to do its part to provide innovative solutions to promote antimicrobial stewardship.
Written by:
Megan Roesner, Clinical Product Manager
Daniel Ballew, Marketing Manager
Kerry Pierce, M.S., RM(NRCM), Design and Development Specialist
Andre Hsiung, M.S., M(ASCP), Business Development Partner
References
- "Core Elements of Antibiotic Stewardship | Antibiotic Use | CDC.” Centers for Disease Control and Prevention, 2019, https://www.cdc.gov/antibiotic-use/core-elements/index.html.
- “Core Elements of Hospital Antibiotic Stewardship Programs | Antibiotic Use | CDC.” Centers for Disease Control and Prevention, 2019, https://www.cdc.gov/antibiotic-use/core-elements/hospital.html#.
- Tan, MD, JD, Siang Yong, and Yvonne Tatsumura, MA, MD. “Alexander Fleming (1881–1955): Discoverer of Penicillin.” PubMed Central (PMC), Singapore Med J, July 2015, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4520913/.
- Rosenblatt-Farrell, Noah. “The Landscape of Antibiotic Resistance.” PubMed Central (PMC), Environment Health Perspectives, June 2009, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2702430/.
- Doron, MD, Shira, and Lisa Davidson, MD. “Antimicrobial Stewardship.” PubMed Central (PMC), Mayo Clinic Proceedings, Nov. 2011, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3203003/#R1.
- “CRE Bacteria: What You Should Know - Mayo Clinic.” Mayo Clinic, Mayo Clinic, 24 Feb. 2021, https://www.mayoclinic.org/diseases-conditions/infectious-diseases/in-depth/cre-bacteria/art-20166387#:~:text=Carbapenem-resistant%20Enterobacteriaceae%20(CRE),cases%20to%20all%20available%20antibiotics.
- Frieden, Thomas R., and Beth P. Bell. “Core Elements of Hospital Antibiotic Stewardship Programs.” American Hospital Association's Physician Leadership Forum, 2014, https://www.ahaphysicianforum.org/resources/appropriate-use/antimicrobial/.
- Healthcare Settings – Preventing the spread of MRSA. Accessed April 21, 2021. https://www.cdc.gov/mrsa/healthcare/index.html
- ESBL-producing Enterobacterales in Healthcare Settings. Accessed April 21, 2021. https://www.cdc.gov/hai/organisms/ESBL.html
- Bradford, P.A. Extended-Spectrum β-Lactamases in the 21st Century: Characterization, Epidemiology, and Detection of this Important Resistance Threat.
- 2001 Oct; 14(4): 933-951. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC89009/pdf/cm0401000933.pdf